The Turkana ecosystem is home to a wide range of species, including many different kinds of insects.
One of the challenges of understanding biodiversity is the fact that many species have not yet been classified, and are in general poorly known or studied. This is true for most of the remote, tropical areas of the world. As a biologist working on insects, it can sometimes seem overwhelming when one is confronted by the incredible diversity of living things that you have to start making sense of. Yet this is a first essential step in asking questions in ecology and evolution.

How do we start to make sense of biodiversity?
In order to properly classify insects, it can be a real challenge as you have to compare specimens from different collections that are spread across the world, as well as work out subtle differences in the morphology and structure of the specimens in front of you. This can take a long time to do.
One solution to this is the use of DNA ‘Barcoding’, which does not replace traditional taxonomy, but works alongside it. This involves extracting DNA and sequencing a gene that scientists have agreed on to use as a benchmark for measuring divergence between sequences, and hence between species and populations. In the case of insects, a mitochondrial gene cytochrome oxidase I (popularly referred to as COI), is widely used as rough species barcode, to help tell things apart. This approach is known as DNA Barcoding and it is enabling scientists to get a rough and ready measure of species diversity that is especially helpful in areas where there is little work done on biodiversity. DNA Barcoding is supported by the Barcode of Life Database initiative and provides a platform for scientists to share information and progress towards building a reference library of sequences for the biodiversity of our planet. Data are freely available to anyone interested and can be accessed through the BOLD website (www.boldsystems.org).
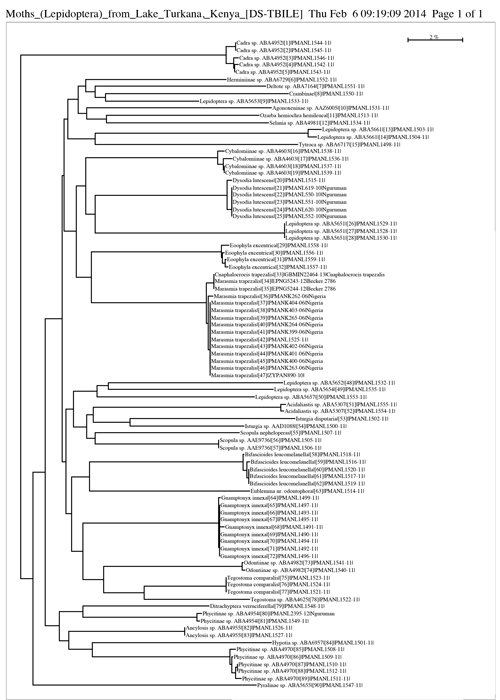
COI Barcode Sequences in tree showing the diversity of the Turkana moths
This useful tool has been applied to a small collection of moths made in Turkana by Dr. Scott Miller of the Smithsonian Institution and myself and sequences generated with the help of Margaret Rosati (Smithsonian Institution) and Dr. Paul D. N. Herbert (University of Guelph, Ontario). This small collection of moths represents the first DNA Barcodes of any form of extant life from the Turkana Basin. This is an important first step in understanding the biodiversity and it’s ecology and evolution in Turkana. In the BOLD system one of the useful tools available are images of the specimens, which can help aid field studies and identifications:
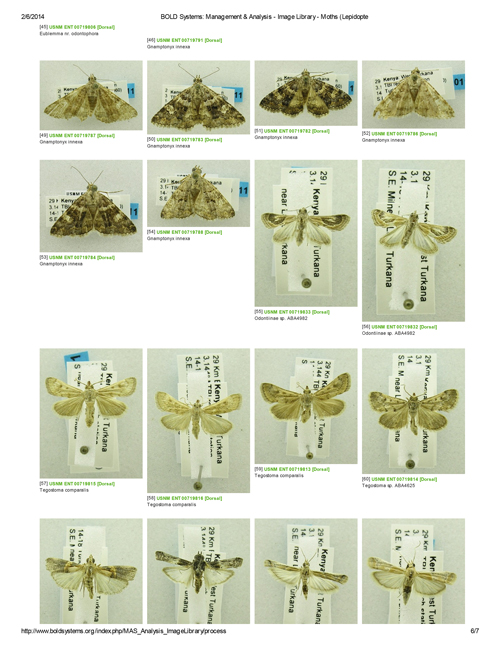

A few of the Turkana moths in the BOLD database
Already the sequence data from the moths of Turkana is being referenced by scientists working in the Middle East and incredibly the COI data show that there is little overlap between the moth fauna of Turkana and other parts of Kenya further south, including Laikipia where a large number of moths have been collected by Dr. Miller. The moth fauna of this small corner of Turkana has a distinctively North African affinity. This work was just published online in the Proceedings of the Entomological Society of Washington.
This is just one of the first scientific steps towards understanding and exploring further the biodiversity of this interesting part of the world.





